From fragments to insights: highlights from SciLake’s closing event
From fragments to insights: highlights from SciLake’s closing event
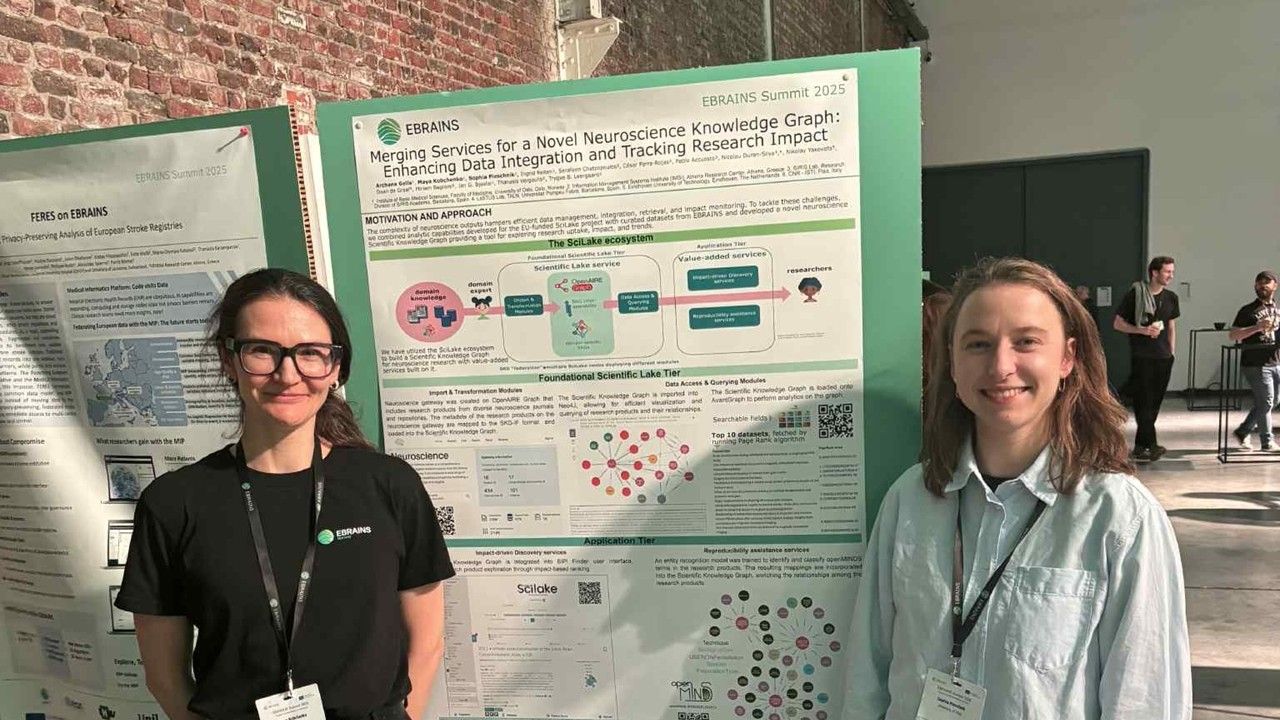
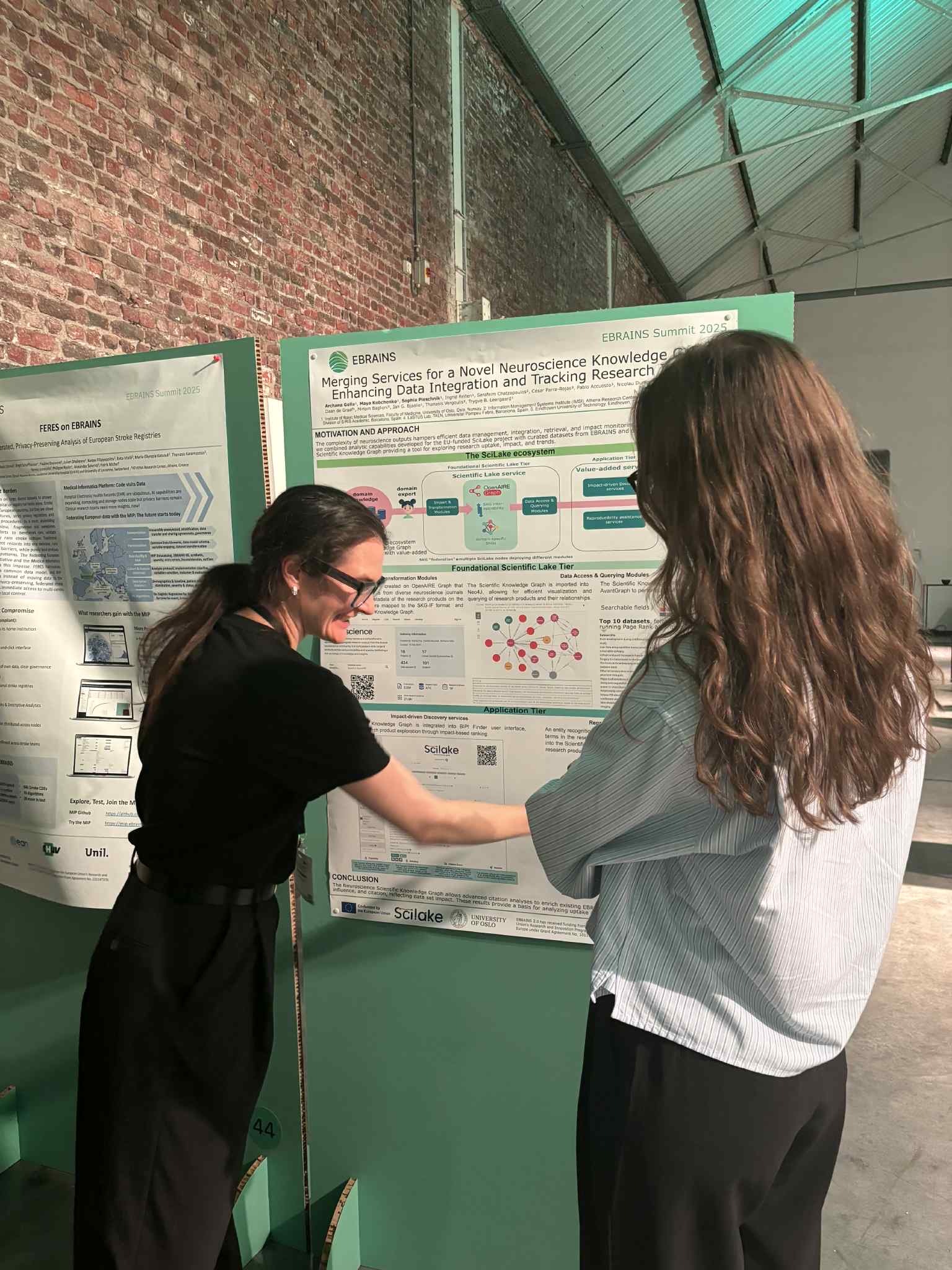
Scientific knowledge is richer than ever, yet hard to navigate. Research outputs are spread across articles, datasets, software, protocols, workflows, and many other sources. Even when content is openly available, it can remain functionally fragmented: difficult to connect, query, reuse, and translate into evidence for new research or policy decisions. Designed to help close this gap, SciLake brought the consortium together for its final dissemination event on Tuesday 10 March 2026, a half‑day online gathering showcasing practical approaches to tackling fragmentation through customisable Scientific Knowledge Graphs (SKGs) piloted in real, domain-specific contexts (neuroscience, cancer, transportation, and energy). This post captures the key takeaways shared across the event.
The SciLake journey and the fragmentation problem
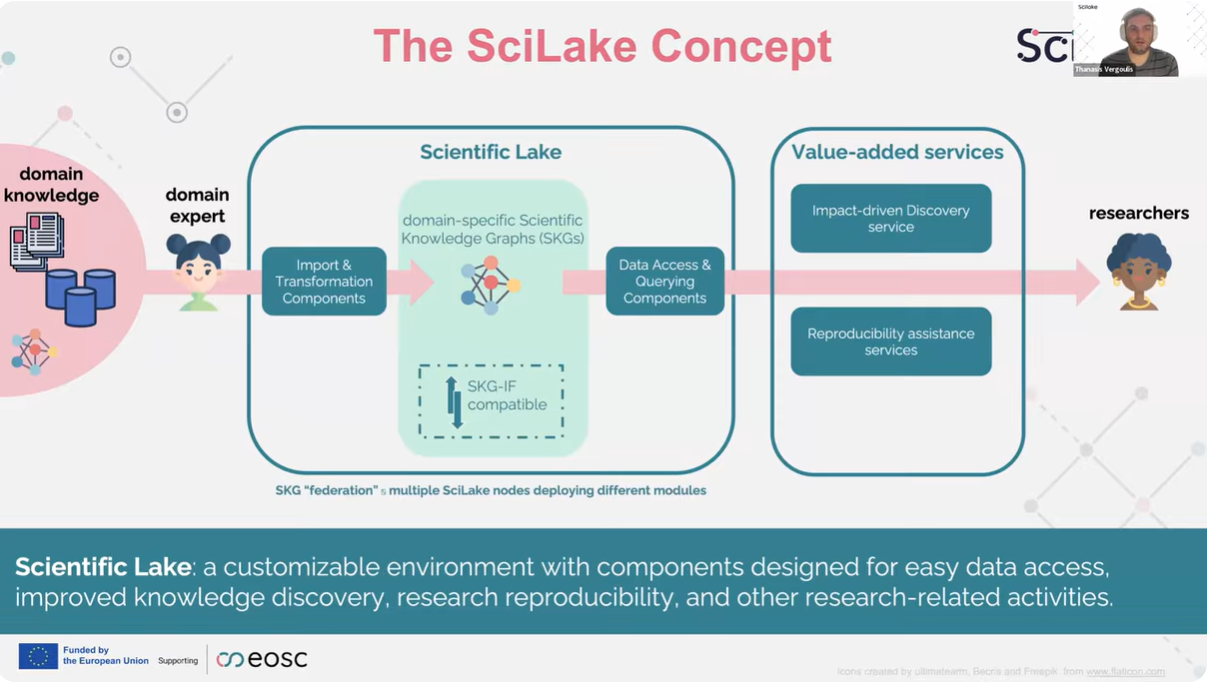
In the opening talk, Thanasis Vergoulis (ARC) framed the challenge: valuable scientific knowledge is abundant, but scattered across heterogeneous sources and encoded in formats that make it hard to connect at scale.
SciLake’s response was presented as a customisable ecosystem of components that helps communities:
- ingest and transform domain corpora,
- generate domain-specific SKGs aligned with standards, especially the RDA Scholarly Interoperability Framework (SKG‑IF)
- expose the graphs via APIs and graph technologies,
- and build value-added services on top.
Two example services illustrated the concept end-to-end:
- a knowledge discovery service using citation-network analysis, and
- a reproducibility assistance service to help identify reproducible studies and flag reproducibility issues.
SciLake’s work is also especially timely in the era of large language models (LLMs) and generative AI: SKGs can provide reliable context and help reduce hallucinations in AI‑assisted discovery and summarisation.
Panel: Turning scattered knowledge into “living” knowledge graphs
Three key themes that emerged from the discussion:
- Build on open backbones, then specialise with communities. A rich, domain-agnostic base like the OpenAIRE Graph can be curated into community-specific subsets through clear criteria and workflows.
- Combine knowledge graphs and large language models (LLMs). SKGs offer traceability, updates, and explicit reasoning; LLMs are strong at conversational access and extraction. Together, LLMs can provide interaction while SKGs provide reliable, auditable knowledge base.
- Make trust practical with provenance and correction loops. Linking research objects is essential, but automated enrichment and linking can introduce errors. Trust therefore depends on being able to trace each link back to its source (provenance) and on having feedback loops so communities can flag issues, correct them, and improve the graph over time.
Pilot demonstrations: SciLake tools through domain case studies
The demo session showed SciLake’s approach in practice across five pilots, each starting from a curated collection and moving toward an interoperable, enriched SKG.
Key highlights across the pilots:
- Energy planning: extracting geographic “objects of study” from publications, linking them to places in OpenStreetMap, and enabling spatial discovery.
- Cancer: a cancer-specific KG integrating ontologies and gene/drug resources, supporting pathway exploration, interaction networks, and literature ranking.
- Maritime transport: stakeholder-driven monitoring and discovery with emphasis on open access and reproducibility. Ranking signals (e.g., influence/PageRank and recency-biased popularity) were computed and released openly.
- CCAM (automated mobility): building targeted subgraphs and reducing noise compared with generic search. Entity extraction and linking for CCAM concepts supported trend spotting, with human-in-the-loop validation.
- Neuroscience: building an SKG enriched with neuroscience entities and impact indicators, enabling discovery and query-based meta-analysis, and helping explore how dataset citation relates to citation impact.
Panel: Unlocking hidden patterns in knowledge graphs
This panel shifted from what is already possible in pilots to the harder questions: what it takes to scale SKGs into sustainable services.
Key takeaways:
- Trust and verification come first. Researchers need to avoid both under-trust (not using tools) and over-trust (uncritical adoption).
- The next step is richer representations. Progress depends on models that support reasoning and synthesis, not only similarity search and summarisation.
- Data ingest remains a bottleneck. PDFs, handwritten tables, and missing abstracts still limit what can become machine actionable.
- Scaling is more than algorithms. It requires sustained investment in infrastructure, interoperability, and long-term sustainability.
Beyond SciLake: takeaways for Open Science
Interoperable SKGs are becoming a practical instrument for Open Science. They help move from “open but scattered” to “open and reusable” by enabling discovery, querying, and evidence-based navigation.
Community knowledge makes the difference. Domain experts are essential for deciding which entities and relationships matter, what “quality” means, and which evidence paths are meaningful.
Provenance and governance are non‑negotiable. Especially when AI is involved, users need to trace claims back to sources, inspect curation decisions, and correct errors.
SciLake’s final event clarified that solving knowledge fragmentation is not about “one more platform.” It is about enabling communities to create and sustain interoperable, trustworthy knowledge structures that evolve with research and that make open research outputs more usable for people, services, and responsible AI.
Read more …From fragments to insights: highlights from SciLake’s closing event